Enzyme
- Define enzyme
- en=in; zyme=yeast
- biological catalyst increase rate of reaction by 1 mil-fold, typically about 1 thousand-fold
- most enzymes are proteins
- PM enzyme→regulate catalysis within cells in response to EC signals
- circulatory enzymes→regulate blood clot
- IC enzyme→catalyze reactions of metabolic pathways
- Define substrate→molecule that binds to an enzyme to yield a product
- Nomenclature
- substrate→lactase catalyze lactose into glucose + galactose
- action→DNase catalyze DNA into dNMP
- Enzyme class 1 main class.1 subclass.1 sub-sub.1 serial no of sub-sub
- oxireductase
- +/- H
- catalyze oxidation-reduction
- 22 subclass
- oxidation of alcohol donor + acceptor NAD+ to aldehyde + NADH→oxidation
- active site→zinc, His-51, 67, Cys-174, 46, Ser-48, IIe-269, Val-292, Ala-317, Phe-319
- transferase
- transfer functional group→amino, acyl, P, glycosyl
- 9 subclass
- ALT→glutamic acid + pyruvic acid to a-ketoglutaric acid + alanine
- hydrolase
- nearly all proteins on outer surface→glycoprotein
- hydrolysis of chemical bond
- breakdown substrate by + H20
- 13 subclass
- serine protease cleave peptide bond
- lyase
- remove H2O NH3 CO2
- cleave C-C, C-O, C-N, C-S, C-H, P-O bond
- 7 subclass
- pyruvate decarboxylase→pyruvate + H to acetaldehyde + CO2
- depends on cofactors TPP and Mg
- 4 active site, each has 20 aa, Glu-477 stability of TPP ring, 51 cofactor binding
- isomerase
- L to D isomerization
- heterogenous group of enzymes catalyze isomerization of cis-trans and aldose-ketose interconversion
- 6 subclass
- D-glucose to trans enediol to D-fructose to cis enediol to D-mannose
- ligase
- 2 chemical group joined with high-energy phosphate bond of ATP is broken→ligate = bind
- 9 subclass
- oxireductase
- Zymogen/proenzyme
- inactive
- trypsin→chymotrypsinogen to chymotrypsin
- enteropeptidase→trypsinogen to trypsin
- prothrombinase→prothrombin to thrombin
- Simple enzyme→proteins + relatively small organic molecule
- Isoenzyme
- catalyze same reaction
- differ structure (aa seq), Km, heat stability, susceptibility to inhibition
- LDH1 in cardiac muscle H4
- LDH2 in RBC H3M
- LDH3 in lungs H2M2
- LDH4 in other tissues HM3
- LDH5 in liver and skeletal muscles M4
- H4 higher affinity for substrate
- heart attack→confirm dimeric CK tetrameric LDH (both 2 protomers) in serum by electrophoresis
- heart muscle→CK1 BB, CK2 MB, CK3 MM
- Multienzyme complex
- catalytic domain→part of (more than one type) polypeptide chain the posses a catalytic function
- >1 structural domain
- a-ketoglutarate dehydrogenase→3 enzymes 5 coenzymes that catalyze oxidative decarboxylation of a-ketoglutarate, coenzyme A, NAD+ to succinyl-CoA, CO2, NADH in TCA
- pyruvate dehydrogenase→3 enzymes 5 cofactors that catalyze oxidative decarboxylation of pyruvate with formation of acetyl-CoA, CO2, NADH
- Holoenzyme=apoenzyme (responsible for reaction) + cofactor (bond formation between enzyme & substrate, transfer functional groups, form tertiary)
- complete, functional enzyme
- DNA polymerase 3, RNA polymerase
- Metal ions
- cofactors→Cu2+, Fe3+, Zn2+
- Cd2+ and Hg2+ can replace Zn2+
- Metalloenzyme
- enzymes require metal ions
- bind and retain metal atoms with very high affinity
- metal-activated enzyme→low affinity for metal ion
- Cofactor
- catalytically essential molecules or ions covalently bound to enzyme
- non-protein chemical compound/metallic ion as catalyst→vitamin, minerals, ATP
- Coenzyme
- organic molecules contain functionalities not found in protein
- nonprotein with low molecular weight and heat-stable
- NAD, NADP, FMN, FAD, lipoic acid, glutathione, coenzyme Q, iron porphyrins (heme), TPP, pyridoxal phosphate, CoA, biotin, tetrahydrofolic acid, ATP
- Cosubstrate
- coenzymes bind tightly to a protein, yet will be released and bind again at some point
- NAD+, NADP+
- alcohol dehydrogenase→ethanol + NAD+ to acetaldehyde + NADH + H
- Prosthetic group
- water soluble vitamin BC
- malfunction of enzyme with lack sufficient cofactors derived from vitamin→dietary vitamin deficiency=functional & dietary
- Specificity
- stereospecificity
- regioselectivity
- chemo selectivity
- 4 process
- Lock key model→active site complementary in conformation to substrate
- Induced fit model→binding site changes shape upon binding with specific substrate
- Active site→pocket or cleft surrounded by aa side chains help bind substrate and by other side chains catalysis occurs
- Serine protease
- reactive serine residue at catalytic site→cleave peptide bond
- chymotrypsin→hydrophobic residue=phe, trp, tyr
- trypsin→positive charge residue/basic residue=lys, arg, his
- elastase→specific for ala, gly, val
- Enzyme kinetic
- rate and mechanism of reaction
- rate measure how many moles of reactant or product changed per time period
- mechanism is how reaction occurs at molecular levels
- rate of reaction depends on concentration of substrate [S]
- rate = v = change in [P]/change in time = k[S]
- rate is directly proportional to [S]
- enzyme convert S to P→forward
- as P grows, back reaction rate increases until equilibrium→reverse
- graph not linear
- as S larger, enzyme becomes limit
- rate depends on fixed enzyme amount
- rate asymptotically approach a max
- rate depends on [S] but hyperbolic curve & plateaus (Vmax) depend on [enzyme]
- Michaelis-Menten model
- E+S to ES to activated ES* to EP to E+P
- km=[S], high km=weak binding
- Michaelis constant→Vmax /2
- Lineweaver burk plot
- Regulatory mechanism
- [S]
- [enzyme]
- [cofactor/coenzyme]
- [inhibitor/activator]
- covalent modification
- temperature→bell curve
- pH
- bell shape
- alkalosis
- acidosis
- time
- compartmentalization
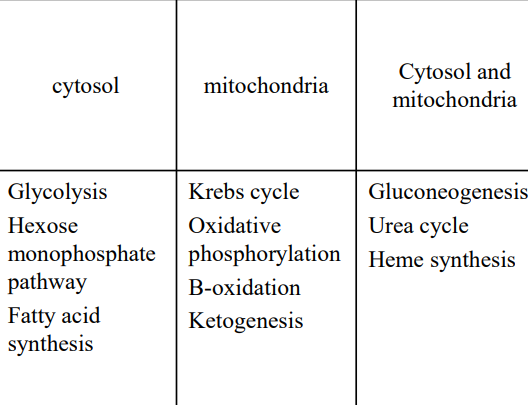
- Inhibitor
- reversible
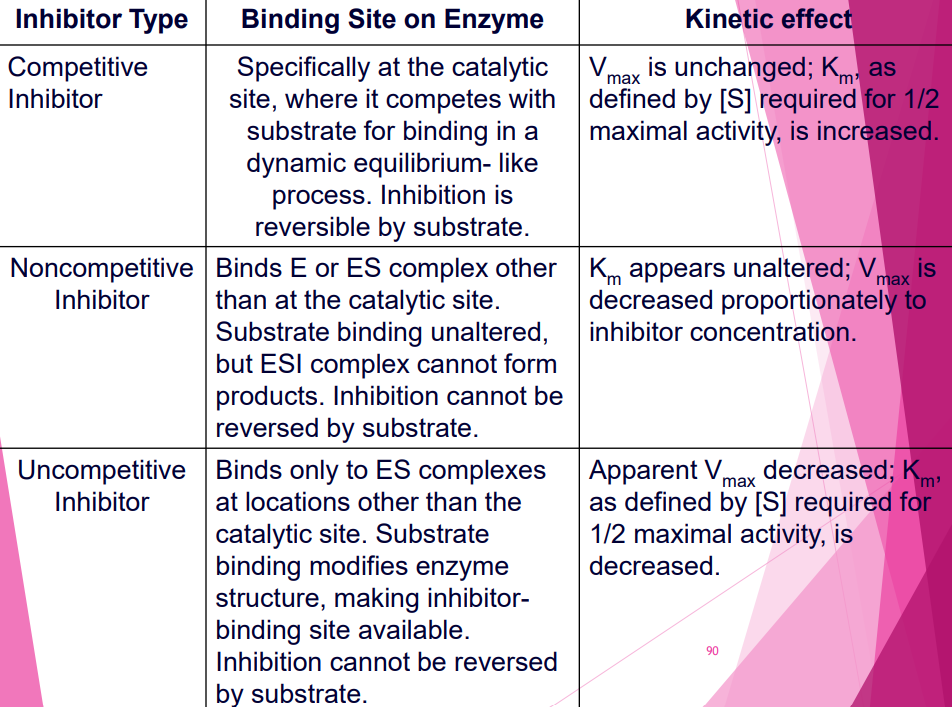
- Competitive inhibitor block active site, Vmax same, high km
- Non-competitive inhibitor + allosteric, alter active site, low Vmax, km same
- Uncompetitive inhibitor + ES, low Vmax, low km
- irreversible
- enzyme inactivator
- E+I=EI
- take time to react with enzyme as slow formation of covalent bonds
- time dependency→degree of inhibition increase with time
- example
- aspirin inhibit COX 1&2
- allopurinol inhibit xanthine oxidase
- ritonavir compete with substrate of HIV protease
- tipranavir non peptide
- methotrexate compete with dihydrofolate and substrate of dihydrofolate reductase, inhibit TMP synthesis, for leukemia
- cyanide irreversible copper/iron in active site of cytochrome c oxidase
- oligomycin inhibit ATP synthase
- S-hexaglutathione
- reversible
- Nerve gas
- diisopropylphosphofluoridate DIPF inhibit Ach ase
- G agents
- sarin
- VX inhibit Ach ase, Na K pump open prolong, muscle contraction prolong


Comments
Post a Comment